 |
| The
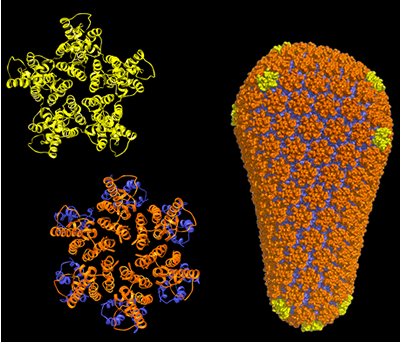
new detailed description of the complete HIV capsid
(outer shell) will provide a roadmap for drugs that
can disrupt its formation and thus prevent infection
by HIV. (Graphics by Owen Pornillos, Barbie Ganser-Pornillos,
Kelly Dryden, and Mark Yeager.) |
|
Below
is the text of a press release issued by the Scripps Research
Institute describing the research and its findings.
Scripps
Research and University of Virginia Scientists Reveal Complete
Structure of HIV's Outer Shell
 |
The
research provides clues for new therapies by shedding light
on how the outer coating of the virus forms |
La Jolla, CA -- January 19, 2011 -- A team of scientists at The
Scripps Research Institute and the University of Virginia has
determined the structure of the protein package that delivers
the genetic material of the human immunodeficiency virus (HIV)
to human cells.
The work is the culmination of studies carried out over the last
decade looking at different portions of the cone-shaped container,
or the capsid. The final piece of the puzzle, described in an
article published in Nature on January 20, 2011, details the structure
of the two ends of the cone.
"This paper is a milestone for research from our group,"
says the study's senior author Mark Yeager, MD, PhD, a Scripps
Research professor and staff cardiologist and chair of the Molecular
Physiology and Biological Physics Department at The University
of Virginia School of Medicine.
A detailed description of the complete HIV capsid will provide
a roadmap for developing drugs that can disrupt its formation
and thus prevent infection by HIV.
Assembling the Package
HIV binds to receptors on human cells and then delivers the capsid
inside them. Once inside a cell, the capsid comes apart, releasing
its precious cargo -- the virus's genetic material.
HIV then sabotages the cell machinery to make many copies of its
genes and proteins. As new viruses are made, the genetic material
is packaged into spherical immature capsids that HIV uses to escape
from the infected cell. But before these newly released viruses
can infect other cells, the immature capsid undergoes a dramatic
rearrangement to form the mature, cone-shaped shell.
If formation of the mature capsid is disrupted, the virus is no
longer infectious. Thus, new drugs targeting capsid formation
could provide valuable additions to the arsenal of existing drugs
against HIV.
A "Floppy" Bridge
To develop drugs that disrupt capsid formation, however, scientists
first need to know precisely how it is formed.
One technology researchers use to obtain detailed structures of
biological molecules is X-ray crystallography. This technique
requires growing crystals of a molecule and then bombarding the
crystals with X-rays to determine the positions of all the atoms.
But unlike the cone-shaped capsids of other viruses, such as the
poliovirus, which have a rigid, symmetrical structure that obediently
assembles into crystals, the HIV capsid is flexible and can adopt
slightly different shapes.
Part of the reason for this flexibility is the protein that makes
up the HIV capsid, the CA protein, consists of two ends held together
by a "floppy" bridge.
The main conical capsid contains about 250 of the six-fold units
or hexamers. Each end of the cone is then closed off by either
five or seven smaller five-fold units or pentamers.
"It is impossible to grow crystals of the entire HIV capsid,"
says Yeager. As a result, his team used a "divide and conquer
approach."
Divide and Conquer
Working primarily with husband-and-wife team Owen Pornillos and
Barbie Ganser-Pornillos, investigators in his lab, Yeager partitioned
the HIV capsid into smaller components, then determined their
respective structures.
In a journey that started more than 10 years ago, Yeager's group
as well as others generated two-dimensional arrays of the hexagonal
CA protein that mimicked the hexagonal lattice of the capsid.
A breakthrough came in 2007, when the Yeager group used electron
microscopy to visualize the CA lattice at higher resolution, which
suggested how the domains of CA are packed together: each CA protein
joins hands with the adjacent CA protein molecules to form the
hexamer.
Guided by the atomic model based on the EM structure, in 2009
the team engineered the CA hexamers so that they formed three-dimensional
crystals suitable for X-ray crystallography. The researchers were
then able to determine the particles' structures at 2-Angstrom
resolution (one Angstrom equals one ten-billionth of a meter).
Having cracked the atomic structure of the hexamer, the investigators
turned their attention to the more elusive pentamers.
Next Came the Pentamer
In this latest study, Yeager, Pornillos, and Ganser-Pornillos
used techniques similar to those they had applied to the hexamers
to obtain the crystal structures of the CA pentamers.
The new structure reveals that five CA proteins link hands at
one end, called the N-terminal domain (NTD), to form a circle.
The opposite ends of the CA proteins, called C-terminal domain
(CTD), form a floppy belt around this central core. Then, CTD
links to CTD to connect adjacent pentamers.
The structure reveals flexibility and mobility both between the
central core and belt within each pentamer and at the CTD-CTD
interfaces of adjacent pentamers. The CTD subunits can rotate
relative NTDs. "As a result, each ring can adopt slightly
different angles relative to its adjacent rings," says Pornillos,
first author of the paper.
The structure of the pentamers is remarkably similar to that of
the hexamers, notes Pornillos, with one important difference.
Because pentamers are smaller than hexamers, the amino acids,
the building blocks of proteins, at the center of the pentamer
ring are closer together than in the hexamer.
Many amino acids have positive or negative charges. When two amino
acids with the same charge are close together they tend to push
each other away. One amino acid in the CA protein, called arginine,
with a positive charge, sits smack in the middle of both the hexamer
and pentamer ring.
Because the arginines in the pentamer are packed much closer together,
they repel one another, making the pentamer a less stable structure
than the hexamer. This may explain why there are many more hexamers
in the mature HIV capsid compared to pentamers.
The only place where pentamers are likely to form is at the capsids'
ends, where the linked CA proteins have to bend dramatically to
close off the capsid-a feat the pentamer is more apt to perform.
"Arginine is the critical switch between hexamer and pentamer
formation," says Yeager. "We can finally explain why
the CA protein would make one or the other."
An Atomic Model of the HIV Capsid
Having solved the atomic structures of both CA hexamers and pentamers,
Yeager and colleagues for the first time were able to build a
complete atomic model of the mature HIV capsid.
The researchers now plan to further refine the model using sophisticated
computer programs to determine the stability of the structure
in different regions and to identify possible "weak"
points they can target using newly designed drugs.
They will also begin studying the structure of the immature capsid
to determine how this version of the capsid transitions to the
mature form-a step in the virus lifecycle that has remained mysterious.
"We don't have the full story yet, but we have volume one,"
says Yeager.
Research for paper "Atomic Level Modeling of the HIV Capsid"
was funded by the U.S. National Institutes of Health and the Center
for the Structural Biology of Host Elements in Egress, Trafficking,
and Assembly of HIV (CHEETAH), which is based at the University
of Utah and directed by Dr. Wesley Sundquist.
About The Scripps
Research Institute
The Scripps Research Institute is one of the world's largest independent,
non-profit biomedical research organizations, at the forefront
of basic biomedical science that seeks to comprehend the most
fundamental processes of life. Scripps Research is internationally
recognized for its discoveries in immunology, molecular and cellular
biology, chemistry, neurosciences, autoimmune, cardiovascular,
and infectious diseases, and synthetic vaccine development. An
institution that evolved from the Scripps Metabolic Clinic founded
by philanthropist Ellen Browning Scripps in 1924, Scripps Research
currently employs approximately 3,000 scientists, postdoctoral
fellows, scientific and other technicians, doctoral degree graduate
students, and administrative and technical support personnel.
Headquartered in La Jolla, California, the institute also includes
Scripps Florida, whose researchers focus on basic biomedical science,
drug discovery, and technology development. Scripps Florida is
located in Jupiter, Florida. For more information, see www.scripps.edu.
Investigator
affiliations: Department of Molecular Physiology and Biological
Physics, University of Virginia School of Medicine, Charlottesville,
VA; Department of Cell Biology, Scripps Research Institute, La
Jolla, CA; Division of Cardiovascular Medicine, Department of
Medicine, University of Virginia Health System, Charlottesville,
VA.
1/28/11
Reference
O Pornillos, BK Ganser-Pornillos, and M Yeager. Atomic-level modelling
of the HIV capsid. Nature 469(7330): 424-427 (Abstract).
January 20, 2011.
Other Source
Scripps
Research Institute. Scripps Research and University of Virginia
Scientists Reveal Complete Structure of HIV's Outer Shell. Press
release. January 19, 2011.
|
